Internationally renowned scientists, Professors Leon Dicks and Willie Perold, have joined forces to create a proof-of-concept nanowire biological sensor (or nanochip) that can identify any pathogen or bacteria, for example, e. coli, Salmonella or cholera in 10 to 15 minutes after being swallowed. The significance of this research is that it can ultimately be utilised to accurately diagnose patients during an epidemic or outbreak using a combination of nanotechnology and microbiology. It also means that “instead of prescribing a broad-based antibiotic, or waiting 48 hours for the lab tests to come back, a doctor will be able to immediately prescribe the correct antibiotic to target the pathogen, and by doing so, put less stress on the body’s immune system.”[1] In layman’s terms it means soon you will be able to swallow a nanochip that can diagnose a bacterial infection faster and more cost-effective than traditional tests including bloodwork and endoscopies. Furthermore, you would be able to isolate specific bacteria to test for. For example, if your concern is water quality, you can set up the sensor to only look for e. coli.
Background
During a discussion between Prof Perold of the Department of Electrical and Electronic Engineering and Prof Dicks of the Department of Science, the two scientists realised that they could join forces and combine their research to develop something unique to meet a specific need in the medical world. At the time, Prof Dicks was developing a gel capsule, which if swallowed, had the potential to diagnose whether a patient had a certain disease or infection. The capsule contained antibodies, and once the capsule was removed it could be established whether the antibodies bonded with the pathogen on the capsule which was the indicator for the presence (or not) of a certain disease or infection. At the same time, Prof Perold’s PhD student, Stanley van den Heever, was working on a nano zinc oxide generator that could generate electricity the moment the nanowires are disturbed. Together they came up with the idea to “combine the sensor with ‘biological bait’ to selectively attract bacteria”. The movement would then generate an electrical signal indicating the presence of the selected bacteria.
How it works
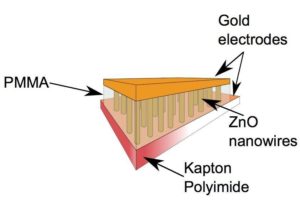
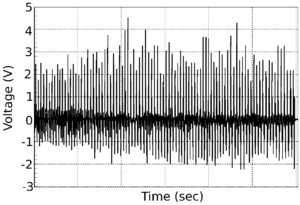
To prepare the nanogenerator for biological use, a silicon chip of 1cm by 1cm had to be constructed. Zinc oxide molecules are subsequently stacked on top of each other to form a nanowire that is only visible with an electron microscope. Thousands of these nanowires are positioned in such a way that the slightest disturbance of their structure will lead to, what is known as, piezoelectric energy. This energy is then converted to electrical energy and amplified to produce a voltage reading.
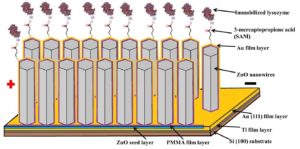
After this process, the microbiologists tested the concept by attaching lysozyme molecules (small disease-fighting proteins found in our saliva) to the tip of each nanowire. As soon as antibodies specific to the lysozyme react to the nanowires, it resulted in an alignment shift of the zinc oxide molecules. This caused a change in electrical output which proved that the concept worked.
If you reverse engineer this concept, it is also possible to attach specific antibodies to the nanowires, which will detect the antigens characteristic of a specific pathogen (that is, a disease-causing microorganism) and report the pathogen presence within seconds. Prof Dicks indicated that the “key to the concept is linking the correct antibody to the nanowires to form a perfect, “one and only”, match with the antigen”.[2]
The future
According to Prof Dicks, their ultimate hope is to transfer the electrical signal explained above, which is selected to be unique to each pathogen, to a receiver such as a smartphone. This means that 1) you can enter remote areas where epidemics or outbreaks are likely to occur and diagnose the disease in minutes as long as you have antibodies available; and 2) it is a more cost effective exercise than, for instance, a endoscopy that requires instruments, a venue and a specialist.
Whether the concept will also be able to identify viruses like Ebola, Prof Dicks is convinced that it will be able to do so as long as you have antibodies specific to the unique Ebola virus antigens. The latter are usually “proteins located on the surface of cells. These proteins act as a ‘signature’ of that specific cell. This ‘signature’ is then recognised by specific antibodies.”[3] This will be a massive step forward in the early diagnosis of Ebola as, traditionally, Ebola is hard to diagnose due to the similarities in its early symptoms to diseases such as malaria, typhoid fever, shigellosis, cholera, leptospirosis and meningitis.
[1] https://blogs.sun.ac.za/microbiology/2014/08/21/nano-chip-in-a-capsule-identifies-bacterial-infections-within-minutes/
[2] https://www.popularmechanics.co.za/science/scientists-develop-sensor-to-identify-disease-causing-bacteria-in-minutes/.
[3] https://www.popularmechanics.co.za/science/scientists-develop-sensor-to-identify-disease-causing-bacteria-in-minutes/.